重测序分析软件安装指南
重测序(resequencing)是指对已知基因组进行高通量测序,以检测个体或种群的遗传变异,从而研究基因组的结构和功能。与全基因组测序不同,重测序通常只对一部分基因组进行测序,例如外显子、基因区域、SNP位点等。
今天分享如何在服务器上部署安装重测序数据分析环境, Linux 服务器下进行重测序分析的常用软件包主要包括:
1. BWA:
用于将测序数据比对到参考基因组上,可以通过 conda 安装:
conda install -c bioconda bwa
2. Samtools:
用于处理 BAM/SAM 格式的测序数据,包括排序、去重、索引等操作,可以通过 conda 安装:
conda install -c bioconda samtools
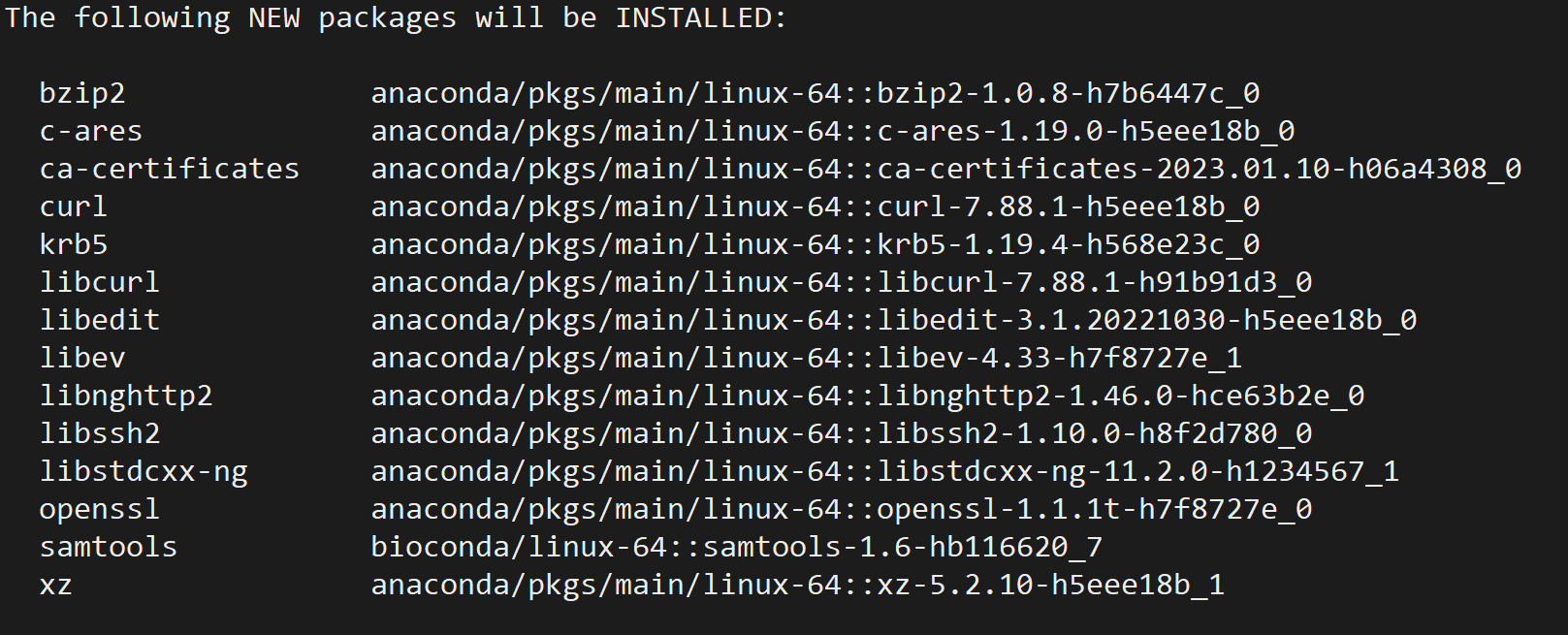
3. Picard:
用于处理 BAM/SAM 格式的测序数据,包括去重、插入大小估计、重命名等操作,可以通过 conda 安装:
conda install -c bioconda picard
4. GATK:
用于进行变异检测和拼接等操作,可以通过 conda 安装:
conda install -c bioconda gatk
5. FastQC:
用于检查测序数据的质量,可以通过 conda 安装:
conda install -c bioconda fastqc
6. Trimmomatic:
用于对测序数据进行质量控制和去除低质量序列,可以通过 conda 安装:
conda install -c bioconda trimmomatic
7. HISAT2:
用于将 RNA-seq 数据比对到基因组上,可以通过 conda 安装:
conda install -c bioconda hisat2
8. StringTie:
用于进行转录本组装和定量分析,可以通过 conda 安装:
conda install -c bioconda stringtie
Linux环境部署结果测试
BWA
(WGS) [root@cloud WGS]# bwa
Program: bwa (alignment via Burrows-Wheeler transformation)
Version: 0.7.17-r1188
Contact: Heng Li <lh3@sanger.ac.uk>
Usage: bwa <command> [options]
Command: index index sequences in the FASTA format
mem BWA-MEM algorithm
fastmap identify super-maximal exact matches
pemerge merge overlapping paired ends (EXPERIMENTAL)
aln gapped/ungapped alignment
samse generate alignment (single ended)
sampe generate alignment (paired ended)
bwasw BWA-SW for long queries
shm manage indices in shared memory
fa2pac convert FASTA to PAC format
pac2bwt generate BWT from PAC
pac2bwtgen alternative algorithm for generating BWT
bwtupdate update .bwt to the new format
bwt2sa generate SA from BWT and Occ
samtools
(WGS) [root@cloud WGS]# samtools
Program: samtools (Tools for alignments in the SAM format)
Version: 1.6 (using htslib 1.6)
Usage: samtools <command> [options]
Commands:
-- Indexing
dict create a sequence dictionary file
faidx index/extract FASTA
index index alignment
-- Editing
calmd recalculate MD/NM tags and '=' bases
fixmate fix mate information
reheader replace BAM header
rmdup remove PCR duplicates
targetcut cut fosmid regions (for fosmid pool only)
addreplacerg adds or replaces RG tags
markdup mark duplicates
通过新建一个conda虚拟环境,然后在环境中安装重测序分析所用的软件,能够避免软件的冲突造成的问题,而且方便再不同的机器之间灵活的切换环境。
重测序可以应用于许多生物学研究领域,如人类疾病研究、植物遗传改良、动物育种等。通过对不同个体或种群的重测序数据进行比较和分析,可以找到与性状相关的遗传变异,揭示基因组的遗传多样性和进化历史,推动生物学研究的发展。
彩蛋
推荐一个重测序分析shell脚本,来自github上某大佬(https://github.com/biomarble/onekeyReseq/blob/main/onekeyReseq),通过这个脚本,能够快捷方便的调用各项软件,自动进行分析,解放双手,提高效率。
echo "$GATKCOMMAND SelectVariants -V $PWD/2.SNP/2.cohort/cohort.vcf -select-type SNP -O $PWD/2.SNP/3.filt/raw.snps.vcf.gz" >commands/2.SNP/s5.Select.sh
echo "$GATKCOMMAND SelectVariants -V $PWD/2.SNP/2.cohort/cohort.vcf -select-type INDEL -O $PWD/2.SNP/3.filt/raw.indel.vcf.gz " >>commands/2.SNP/s5.Select.sh
echo "$GATKCOMMAND VariantFiltration -V $PWD/2.SNP/3.filt/raw.snps.vcf.gz -O $PWD/2.SNP/3.filt/snps.vcf.gz -filter \"QD < 2.0\" --filter-name \"QD2\" -filter \"QUAL < 30.0\" --filter-name \"QUAL30\" -filter \"SOR > 3.0\" --filter-name \"SOR3\" -filter \"FS > 60.0\" --filter-name \"FS60\" -filter \"MQ < 40.0\" --filter-name \"MQ40\" -filter \"MQRankSum < -12.5\" --filter-name \"MQRankSum-12.5\" -filter \"ReadPosRankSum < -8.0\" --filter-name \"ReadPosRankSum-8\" " >commands/2.SNP/s6.Filter.sh
echo "$GATKCOMMAND VariantFiltration -V $PWD/2.SNP/3.filt/raw.indel.vcf.gz -O $PWD/2.SNP/3.filt/indel.vcf.gz -filter \"QD < 2.0\" --filter-name \"QD2\" -filter \"QUAL < 30.0\" --filter-name \"QUAL30\" -filter \"FS > 200.0\" --filter-name \"FS200\" -filter \"ReadPosRankSum < -20.0\" --filter-name \"ReadPosRankSum-20\" " >>commands/2.SNP/s6.Filter.sh
echo "vcftools --gzvcf $PWD/2.SNP/3.filt/snps.vcf.gz --out $PWD/2.SNP/3.filt/final.snp --recode --remove-filtered-all " >commands/2.SNP/s7.extract.sh
echo "vcftools --gzvcf $PWD/2.SNP/3.filt/indel.vcf.gz --out $PWD/2.SNP/3.filt/final.indel --recode --remove-filtered-all " >>commands/2.SNP/s7.extract.sh
echo "snpEff eff -dataDir $PWD -configOption genome.genome=\"genome\" genome $PWD/2.SNP/3.filt/final.snp.recode.vcf -ud 0 -noLog -noStats -geneId -o gatk >$PWD/2.SNP/snp.anno.vcf ">commands/2.SNP/s8.anno.sh
echo "snpEff eff -dataDir $PWD -configOption genome.genome=\"genome\" genome $PWD/2.SNP/3.filt/final.indel.recode.vcf -ud 0 -noLog -noStats -geneId -o gatk >$PWD/2.SNP/indel.anno.vcf" >>commands/2.SNP/s8.anno.sh
echo "$GATKCOMMAND MergeVcfs -I $PWD/2.SNP/indel.anno.vcf -I $PWD/2.SNP/snp.anno.vcf -O $PWD/2.SNP/snp.indel.anno.vcf && bcftools annotate --set-id +'%CHROM\_%POS' $PWD/2.SNP/snp.indel.anno.vcf >$PWD/2.SNP/snp.indel.anno.addid.vcf " >commands/2.SNP/s9.merge.sh
parallelrun "commands/1.mapping/s3.MarkDup.sh"
parallelrun "commands/2.SNP/s1.HC.sh"
parallelrun "commands/2.SNP/s2.CombineGVCFs.sh"
parallelrun "commands/2.SNP/s3.GenotypeGVCF.sh"
parallelrun "commands/2.SNP/s4.CombineVCFs.sh"
parallelrun "commands/2.SNP/s5.Select.sh"
parallelrun "commands/2.SNP/s6.Filter.sh"
parallelrun "commands/2.SNP/s7.extract.sh"
parallelrun "commands/2.SNP/s8.anno.sh"
本文由 mdnice 多平台发布