文章目录
- 安装
- GTDB-Tk v2.3.3 and gtdbtk_r214_data.tar.gz
- GTDB-Tk v2.1.1 and gtdbtk_r207_v2_data.tar.gz
- GTDB-Tk 1.3.0
- 数据库
- classify_wf 物种注释一步使用
- 分步使用
- 批量dRep以及GTDB注释
- 注意
- 报错
- 由于基因组存在重复id导致Pfam报错
- 查看已经安装的数据库
- iTOL可视化问题
- pplacer只是插入树的问题
- 问题
- 参考
安装
GTDB-Tk v2.3.3 and gtdbtk_r214_data.tar.gz
https://data.gtdb.ecogenomic.org/releases/release214/214.0/auxillary_files/
(gtdbtk-2.1.1) [yut@node01 02_Glacier_new_taxa]$ conda env config vars set GTDBTK_DATA_PATH="/datanode02/yut/Database/GTDBTK_r214_data/" # 配置数据库环境变量
To make your changes take effect please reactivate your environment
GTDB-Tk v2.1.1 and gtdbtk_r207_v2_data.tar.gz
Executing transaction: /
GTDB-Tk v2.1.1 requires ~63G of external data which needs to be downloaded
and extracted. This can be done automatically, or manually.
Automatic:
1. Run the command "download-db.sh" to automatically download and extract to:
/datanode02/yut/Software/Miniconda3/envs/gtdbtk-2.1.1/share/gtdbtk-2.1.1/db/
Manual:
1. Manually download the latest reference data:
wget https://data.gtdb.ecogenomic.org/releases/release207/207.0/auxillary_files/gtdbtk_r207_v2_data.tar.gz
2. Extract the archive to a target directory:
tar -xvzf gtdbtk_r207_v2_data.tar.gz -C "/path/to/target/db" --strip 1 > /dev/null
rm gtdbtk_r207_v2_data.tar.gz
3. Set the GTDBTK_DATA_PATH environment variable by running:
conda env config vars set GTDBTK_DATA_PATH="/path/to/target/db"
- 问题
EXCEPTION: ProdigalException
MESSAGE: An exception was caught while running Prodigal: Prodigal returned a non-zero exit code.
AttributeError: module 'numpy' has no attribute 'bool'.
解决方法:
(gtdbtk-2.1.1) [yut@node01 02_Glacier_new_taxa]$ mamba install numpy=1.23.1
GTDB-Tk 1.3.0
conda install -c bioconda gtdbtk
GTDB-Tk v1.3.0 requires ~25G of external data which needs to be downloaded
and unarchived. This can be done automatically, or manually:
1. Run the command download-db.sh to automatically download to:
/home/yutao/miniconda3/share/gtdbtk-1.3.0/db/
2. Manually download the latest reference data:
https://github.com/Ecogenomics/GTDBTk#gtdb-tk-reference-data
2b. Set the GTDBTK_DATA_PATH environment variable in the file:
/home/yutao/miniconda3/etc/conda/activate.d
(gtdbtk) [yutao@myosin GTDB_R202]$ vi ~/miniconda3/envs/gtdbtk/etc/conda/activate.d/gtdbtk.sh
(gtdbtk) [yutao@myosin GTDB_R202]$ cat ~/miniconda3/envs/gtdbtk/etc/conda/activate.d/gtdbtk.sh
export GTDBTK_DATA_PATH=/share/pasteur/yutao/DATABASE/GTDB_R202/release202/
#export GTDBTK_DATA_PATH=/home/yutao/miniconda3/envs/gtdbtk/share/gtdbtk-1.7.0/db/
数据库
wget https://data.ace.uq.edu.au/public/gtdb/data/releases/release95/95.0/auxillary_files/gtdbtk_r95_data.tar.gz
tar xvzf gtdbtk_r95_data.tar.gz
#GTDB-Tk requires an environment variable named GTDBTK_DATA_PATH to be set to the directory containing the unarchived GTDB-Tk reference data.
export GTDBTK_DATA_PATH=/path/to/release/package/
# 最新版R202
https://data.gtdb.ecogenomic.org/releases/release202/202.0/auxillary_files/gtdbtk_r202_data.tar.gz
# 参考
https://ecogenomics.github.io/GTDBTk/installing/index.html
推荐使用flash download下载到本地,在通过filezila上传到服务器
classify_wf 物种注释一步使用
# 输入基因组目录
gtdbtk classify_wf --cpus 20 --genome_dir HTR9_maxbin_genome -x fasta --out_dir classify
# 输入基因组路径,第一列是基因组路径,第二列是基因组名称,便于在后面结果显示,一二列必选;第三列是translation table可选
(gtdbtk-2.1.1) [yut@io02 TestData]$ jobs
[1]+ 运行中 nohup time gtdbtk classify_wf --batchfile test_genome.path --out_dir gtdb_out --cpus 10 &>gtdb.log &
(gtdbtk-2.1.1) [yut@io02 TestData]$ head test_genome.path
/datanode02/yut/Software/TestData/COVID19.fa COVID19.fa
/datanode02/yut/Software/TestData/DRR331386_ma_bin.12.fa DRR331386_ma_bin.12.fa
/datanode02/yut/Software/TestData/DRR331386_ma_bin.18.fa DRR331386_ma_bin.18.fa
–genome_dir 包含多个基因组的目录
-x 基组文件后缀名
- 注意:GTDB-Tk v0.3.2 线程不要超过30个,否则pplacer会一直卡在注释古菌步骤
- 避免滤掉基因组
- 主要结果:物种注释表
(gtdbtk-2.1.1) [yut@io02 TestData]$ cat gtdb_out/gtdbtk.bac120.summary.tsv
user_genome classification fastani_reference fastani_reference_radius fastani_taxonomy fastani_ani fastani_af closest_placement_reference closest_placement_radius closest_placement_taxonomy closest_placement_ani closest_placement_af pplacer_taxonomy classification_method note other_related_references(genome_id,species_name,radius,ANI,AF) msa_percent translation_table red_value warnings
COVID19.fa Unclassified N/A N/A N/A N/A N/A N/A N/A N/A N/A N/A N/A N/A N/A N/A N/A N/A N/A No bacterial or archaeal marker
DRR331386_ma_bin.12.fa d__Bacteria;p__Proteobacteria;c__Gammaproteobacteria;o__Burkholderiales;f__Burkholderiaceae;g__Rhodoferax;s__ N/A N/A N/A N/A N/A N/A N/A N/A N/A N/A d__Bacteria;p__Proteobacteria;c__Gammaproteobacteria;o__Burkholderiales;f__Burkholderiaceae;g__Rhodoferax;s__ taxonomic classification defined by topology and ANI classification based on placement in class-level tree GCA_010365055.1, s__Rhodoferax sp010365055, 95.0, 86.38, 0.64; GCF_000013605.1, s__Rhodoferax ferrireducens, 95.0, 86.12, 0.68; GCA_013791765.1, s__Rhodoferax sp013791765, 95.0, 85.54, 0.65; GCA_002413825.1, s__Rhodoferax sp002413825, 95.0, 80.4, 0.48; GCA_012927445.1, s__Rhodoferax sp012927445, 95.0, 80.38, 0.47; GCF_003347515.1, s__Rhodoferax sp003347515, 95.0, 80.21, 0.5; GCA_017880985.1, s__Rhodoferax sp017880985, 95.0, 80.07, 0.46; GCA_002422455.1, s__Rhodoferax sp002422455, 95.0, 79.42, 0.47; GCA_903828665.1, s__Rhodoferax sp903828665, 95.0, 79.26, 0.38; GCA_002842265.1, s__Rhodoferax sp002842265, 95.0, 79.26, 0.37; GCA_014859475.1, s__Rhodoferax sp014859475, 95.0, 79.23, 0.42; GCA_903839085.1, s__Rhodoferax sp903839085, 95.0, 79.18, 0.33; GCA_011391645.1, s__Rhodoferax sp011391645, 95.0, 79.12, 0.41; GCA_903876735.1, s__Rhodoferax sp903876735, 95.0, 79.09, 0.39; GCA_903920695.1, s__Rhodoferax sp903920695, 95.0, 78.98, 0.39; GCA_011391545.1, s__Rhodoferax sp011391545, 95.0, 78.98, 0.34; GCA_903914175.1, s__Rhodoferax sp903914175, 95.0, 78.95, 0.35; GCA_018780565.1, s__Rhodoferax sp018780565, 95.0, 78.94, 0.39; GCA_002789915.1, s__Rhodoferax sp002789915, 95.0, 78.86, 0.38; GCA_003133245.1, s__Rhodoferax sp003133245, 95.0, 78.84, 0.37; GCA_016708345.1, s__Rhodoferax sp016708345, 95.0, 78.81, 0.36; GCA_903905735.1, s__Rhodoferax sp903905735, 95.0, 78.81, 0.36; GCA_007280205.1, s__Rhodoferax sp007280205, 95.0, 78.8, 0.28; GCA_002083775.1, s__Rhodoferax ferrireducens_A, 95.0, 78.78, 0.37; GCA_903938955.1, s__Rhodoferax sp903938955, 95.0, 78.76, 0.29; GCA_001770935.1, s__Rhodoferax sp001770935, 95.0, 78.66, 0.35; GCA_903960915.1, s__Rhodoferax sp903960915, 95.0, 78.66, 0.32; GCA_903863395.1, s__Rhodoferax sp903863395, 95.0, 78.63, 0.32; GCA_001771065.1, s__Rhodoferax sp001771065, 95.0, 78.6, 0.32; GCF_001938565.1, s__Rhodoferax antarcticus, 95.0, 78.6, 0.32; GCA_903896425.1, s__Rhodoferax sp903896425, 95.0, 78.6, 0.3; GCF_018619435.1, s__Rhodoferax sp018619435, 95.0, 78.58, 0.34; GCA_002786915.1, s__Rhodoferax sp002786915, 95.0, 78.55, 0.35; GCF_013416775.1, s__Rhodoferax sp013416775, 95.0, 78.53, 0.34; GCA_002784245.1, s__Rhodoferax sp002784245, 95.0, 78.52, 0.4; GCA_016718155.1, s__Rhodoferax sp016718155, 95.0, 78.51, 0.35; GCF_002017865.1, s__Rhodoferax fermentans, 95.0, 78.5, 0.34; GCA_903823025.1, s__Rhodoferax sp903823025, 95.0, 78.47, 0.31; GCA_903873485.1, s__Rhodoferax sp903873485, 95.0, 78.47, 0.35; GCA_903897005.1, s__Rhodoferax sp903897005, 95.0, 78.4, 0.28; GCF_001955715.1, s__Rhodoferax saidenbachensis, 95.0, 78.36, 0.3; GCA_001795455.1, s__Rhodoferax sp001795455, 95.0, 78.31, 0.32; GCF_003415675.1, s__Rhodoferax sp003415675, 95.0, 78.3, 0.35; GCA_001871515.1, s__Rhodoferax sp001871515, 95.0, 78.3, 0.32; GCA_008080595.1, s__Rhodoferax sp008080595, 95.0, 78.3, 0.32; GCA_016703945.1, s__Rhodoferax sp016703945, 95.0, 78.26, 0.32; GCA_903894475.1, s__Rhodoferax sp903894475, 95.0, 78.23, 0.3; GCA_009693325.1, s__Rhodoferax sp009693325, 95.0, 78.19, 0.25; GCA_903845935.1, s__Rhodoferax sp903845935, 95.0, 78.15, 0.3; GCA_016718405.1, s__Rhodoferax sp016718405, 95.0, 78.14, 0.33; GCA_008015235.1, s__Rhodoferax sp008015235, 95.0, 78.13, 0.28; GCA_016719055.1, s__Rhodoferax sp016719055, 95.0, 78.08, 0.33; GCF_002943465.1, s__Rhodoferax sp002943465, 95.0, 78.07, 0.32; GCA_014190675.1, s__Rhodoferax sp014190675, 95.0, 78.06, 0.23; GCA_016719405.1, s__Rhodoferax sp016719405, 95.0, 78.05, 0.28; GCA_903897925.1, s__Rhodoferax sp903897925, 95.0, 78.04, 0.27; GCF_017948265.1, s__Rhodoferax sp017948265, 95.0, 78.02, 0.28; GCF_017798165.1, s__Rhodoferax sp017798165, 95.0, 78.01, 0.31; GCA_903853895.1, s__Rhodoferax sp903853895, 95.0, 78.01, 0.26; GCF_002251855.1, s__Rhodoferax sp002251855, 95.0, 77.99, 0.28; GCA_018063565.1, s__Rhodoferax sp018063565, 95.0, 77.97, 0.26; GCA_903886165.1, s__Rhodoferax sp903886165, 95.0, 77.97, 0.32; GCA_903840825.1, s__Rhodoferax sp903840825, 95.0, 77.95, 0.24; GCA_903906695.1, s__Rhodoferax sp903906695, 95.0, 77.95, 0.32; GCA_016705575.1, s__Rhodoferax sp016705575, 95.0, 77.9, 0.28; GCA_903927295.1, s__Rhodoferax sp903927295, 95.0, 77.85, 0.27; GCA_009925925.1, s__Rhodoferax sp009925925, 95.0, 77.84, 0.21; GCA_018061265.1, s__Rhodoferax sp018061265, 95.0, 77.83, 0.22; GCA_903928635.1, s__Rhodoferax sp903928635, 95.0, 77.83, 0.27; GCA_002256115.1, s__Rhodoferax sp002256115, 95.0, 77.82, 0.26; GCA_903945325.1, s__Rhodoferax sp903945325, 95.0, 77.81, 0.27; GCA_017997975.1, s__Rhodoferax sp017997975, 95.0, 77.79, 0.23; GCA_013297935.1, s__Rhodoferax sp013297935, 95.0, 77.79, 0.23; GCA_903864505.1, s__Rhodoferax sp903864505, 95.0, 77.78, 0.2; GCA_009922295.1, s__Rhodoferax sp009922295, 95.0, 77.78, 0.21; GCF_005876985.1, s__Rhodoferax bucti, 95.0, 77.77, 0.29; GCF_002163715.1, s__Rhodoferax sp002163715, 95.0, 77.77, 0.23; GCA_903925725.1, s__Rhodoferax sp903925725, 95.0, 77.7, 0.21; GCA_903944555.1, s__Rhodoferax sp903944555, 95.0, 77.64, 0.18; GCA_903948045.1, s__Rhodoferax sp903948045, 95.0, 77.61, 0.24; GCA_903856025.1, s__Rhodoferax sp903856025, 95.0, 77.61, 0.25; GCA_903930255.1, s__Rhodoferax sp903930255, 95.0, 77.57, 0.19; GCA_903924955.1, s__Rhodoferax sp903924955, 95.0, 77.54, 0.24; GCA_903876045.1, s__Rhodoferax sp903876045, 95.0, 77.49, 0.21; GCA_018882485.1, s__Rhodoferax sp018882485, 95.0, 77.47, 0.22; GCA_903954045.1, s__Rhodoferax sp903954045, 95.0, 77.45, 0.2; GCA_903958915.1, s__Rhodoferax sp903958915, 95.0, 77.43, 0.19; GCA_903869595.1, s__Rhodoferax sp903869595, 95.0, 77.43, 0.21; GCA_009924455.1, s__Rhodoferax sp009924455, 95.0, 77.41, 0.23; GCA_903875275.1, s__Rhodoferax sp903875275, 95.0, 77.34, 0.23; GCA_903859265.1, s__Rhodoferax sp903859265, 95.0, 77.32, 0.19; GCA_903928325.1, s__Rhodoferax sp903928325, 95.0, 77.31, 0.21; GCA_002390825.1, s__Rhodoferax sp002390825, 95.0, 77.3, 0.19; GCF_006974105.1, s__Rhodoferax sp006974105, 95.0, 77.2, 0.19; GCA_002381045.1, s__Rhodoferax sp002381045, 95.0, 77.19, 0.22; GCA_903870955.1, s__Rhodoferax sp903870955, 95.0, 77.02, 0.18; GCA_007279715.1, s__Rhodoferax sp007279715, 95.0, 77.01, 0.11; GCA_903914985.1, s__Rhodoferax sp903914985, 95.0, 76.98, 0.22; GCA_903870065.1, s__Rhodoferax sp903870065, 95.0, 76.94, 0.19; GCA_005793295.1, s__Rhodoferax sp005793295, 95.0, 75.99, 0.1 81.04 11 0.9887676016165975 Genome not assigned to closest species as it falls outside its pre-defined ANI radius;Genome has more than 11.7% of markers with multiple hits
DRR331386_ma_bin.18.fa d__Bacteria;p__Patescibacteria;c__Saccharimonadia;o__UBA4664;f__UBA4664;g__UBA4664;s__ N/A N/A N/A N/A N/A GCA_002415345.1 95.0 d__Bacteria;p__Patescibacteria;c__Saccharimonadia;o__UBA4664;f__UBA4664;g__UBA4664;s__UBA4664 sp00241534582.82 0.8 d__Bacteria;p__Patescibacteria;c__Saccharimonadia;o__UBA4664;f__UBA4664;g__UBA4664;s__ taxonomic classification defined by topology and ANI classification based on placement in class-level tree N/A 65.55 11 0.9673951391491065 Genome not assigned to closest species as it falls outside its pre-defined ANI radius
- 时间:3179个基因组28核耗时30h
- 标记基因蛋白多序列文件及其他结果
(gtdbtk-2.1.1) [yut@io02 TestData]$ tree gtdb_out
gtdb_out
├── align
│ ├── gtdbtk.bac120.filtered.tsv
│ ├── gtdbtk.bac120.msa.fasta.gz # gtdb所有物种细菌120个标记
│ └── gtdbtk.bac120.user_msa.fasta.gz # 输入基因组细菌120个标记
├── classify
│ ├── gtdbtk.bac120.classify.tree.1.tree
│ ├── gtdbtk.bac120.classify.tree.6.tree
│ ├── gtdbtk.bac120.summary.tsv
│ ├── gtdbtk.bac120.tree.mapping.tsv
│ └── gtdbtk.backbone.bac120.classify.tree
├── gtdbtk.bac120.summary.tsv -> classify/gtdbtk.bac120.summary.tsv
├── gtdbtk.log
├── gtdbtk.warnings.log
└── identify
├── gtdbtk.ar53.markers_summary.tsv
├── gtdbtk.bac120.markers_summary.tsv
├── gtdbtk.failed_genomes.tsv
└── gtdbtk.translation_table_summary.tsv
分步使用
- 原始数据
原始数据为基因组,例如Genome A:GCF_003947435.1 [GTDB ] * Genome B: GCA_002011125.1 [GTDB]
# Create the directory.
mkdir -p /tmp/gtdbtk && cd /tmp/gtdbtk
# Obtain the genomes.
mkdir -p /tmp/gtdbtk/genomes
wget -q https://ftp.ncbi.nlm.nih.gov/genomes/all/GCF/003/947/435/GCF_003947435.1_ASM394743v1/GCF_003947435.1_ASM394743v1_genomic.fna.gz -O /tmp/gtdbtk/genomes/genome_a.fna.gz
wget -q https://ftp.ncbi.nlm.nih.gov/genomes/all/GCA/002/011/125/GCA_002011125.1_ASM201112v1/GCA_002011125.1_ASM201112v1_genomic.fna.gz -O /tmp/gtdbtk/genomes/genome_b.fna.gz
- Step 2: Gene calling (identify)
可以通过单步完成: classify_wf,但是分部完成更适合处理大数据
ls -l /tmp/gtdbtk/genomes
total 1464
-rw-rw-r–. 1 uqamussi uqamussi 877141 Mar 12 2019 genome_a.fna.gz
-rw-rw-r–. 1 uqamussi uqamussi 616683 Mar 3 2017 genome_b.fna.gz
gtdbtk identify --genome_dir /tmp/gtdbtk/genomes --out_dir /tmp/gtdbtk/identify --extension gz --cpus 2
–genome_dir,包含多个基因组的目录,例如binning后生成的多个bins
–extension,是基因组文件的后缀名
[2020-08-03 11:03:42] INFO: GTDB-Tk v1.3.0
[2020-08-03 11:03:42] INFO: gtdbtk identify --genome_dir /tmp/gtdbtk/genomes --out_dir /tmp/gtdbtk/identify --extension gz --cpus 2
[2020-08-03 11:03:42] INFO: Using GTDB-Tk reference data version r95: /srv/db/gtdbtk/official/release95
[2020-08-03 11:03:42] INFO: Identifying markers in 2 genomes with 2 threads.
[2020-08-03 11:03:42] INFO: Running Prodigal V2.6.3 to identify genes.
==> Processed 2/2 (100%) genomes [ 6.28s/it, ETA 00:00]
[2020-08-03 11:03:54] INFO: Identifying TIGRFAM protein families.
==> Processed 2/2 (100%) genomes [ 3.79s/it, ETA 00:00]
[2020-08-03 11:04:02] INFO: Identifying Pfam protein families.
==> Processed 2/2 (100%) genomes [ 1.62it/s, ETA 00:00]
[2020-08-03 11:04:03] INFO: Annotations done using HMMER 3.1b2 (February 2015).
[2020-08-03 11:04:03] INFO: Done.
- Results
可以在每个基因组对应的中间文件目录下找到被调用的基因和标记信息,如下所示。
ls /tmp/gtdbtk/identify/identify/intermediate_results/marker_genes/genome_a.fna/
genome_a.fna_pfam_tophit.tsv genome_a.fna_protein.gff.sha256
genome_a.fna_pfam_tophit.tsv.sha256 genome_a.fna_tigrfam.out
genome_a.fna_pfam.tsv genome_a.fna_tigrfam.out.sha256
genome_a.fna_pfam.tsv.sha256 genome_a.fna_tigrfam_tophit.tsv
genome_a.fna_protein.faa genome_a.fna_tigrfam_tophit.tsv.sha256
genome_a.fna_protein.faa.sha256 genome_a.fna_tigrfam.tsv
genome_a.fna_protein.fna genome_a.fna_tigrfam.tsv.sha256
genome_a.fna_protein.fna.sha256 prodigal_translation_table.tsv
genome_a.fna_protein.gff prodigal_translation_table.tsv.sha256
然而,有时仅仅阅读概要文件更有用,该概要文件详细描述了从古细菌122或细菌120标记集中识别的标记。 (archaeal 122, or bacterial 120 marker set.)
cat /tmp/gtdbtk/identify/gtdbtk.ar122.markers_summary.tsv
name number_unique_genes number_multiple_genes number_multiple_unique_genes number_missing_genes list_unique_genes list_multiple_genes list_multiple_unique_genes list_missing_genes
genome_a.fna 109 3 0 10 PF00368.13,PF00410.14,PF00466.15,PF00687.16,PF00827.12,PF00900.15,PF01000.21,PF01015.13,PF01090.14,PF01092.14,PF01157.13,PF01191.14,PF01194.12,PF01198.14,PF01200.13,PF01269.12,PF01280.15,PF01282.14,PF01655.13,PF01798.13,PF01864.12,PF01868.11,PF01984.15,PF01990.12,PF02006.11,PF02978.14,PF03874.11,PF04019.7,PF04104.9,PF04919.7,PF07541.7,PF13656.1,PF13685.1,TIGR00021,TIGR00037,TIGR00042,TIGR00111,TIGR00134,TIGR00240,TIGR00264,TIGR00270,TIGR00279,TIGR00283,TIGR00291,TIGR00293,TIGR00307,TIGR00308,TIGR00323,TIGR00324,TIGR00335,TIGR00336,TIGR00337,TIGR00389,TIGR00392,TIGR00398,TIGR00405,TIGR00408,TIGR00422,TIGR00425,TIGR00432,TIGR00442,TIGR00448,TIGR00456,TIGR00463,TIGR00468,TIGR00471,TIGR00490,TIGR00491,TIGR00501,TIGR00521,TIGR00549,TIGR00670,TIGR00729,TIGR00936,TIGR00982,TIGR01008,TIGR01012,TIGR01018,TIGR01020,TIGR01025,TIGR01028,TIGR01038,TIGR01046,TIGR01052,TIGR01060,TIGR01077,TIGR01080,TIGR01213,TIGR01309,TIGR01952,TIGR02076,TIGR02153,TIGR02236,TIGR02258,TIGR02390,TIGR02651,TIGR03626,TIGR03627,TIGR03628,TIGR03636,TIGR03653,TIGR03665,TIGR03671,TIGR03672,TIGR03674,TIGR03677,TIGR03680,TIGR03683,TIGR03684 PF01496.14,TIGR00458,TIGR00658 PF01866.12,TIGR00064,TIGR00373,TIGR00522,TIGR02338,TIGR02389,TIGR03629,TIGR03670,TIGR03673,TIGR03722
genome_b.fna 118 2 0 2 PF00368.13,PF00410.14,PF00466.15,PF00687.16,PF00827.12,PF00900.15,PF01000.21,PF01015.13,PF01090.14,PF01092.14,PF01157.13,PF01191.14,PF01194.12,PF01198.14,PF01200.13,PF01269.12,PF01280.15,PF01282.14,PF01655.13,PF01798.13,PF01864.12,PF01866.12,PF01868.11,PF01984.15,PF01990.12,PF02006.11,PF02978.14,PF03874.11,PF04019.7,PF04104.9,PF04919.7,PF07541.7,PF13656.1,TIGR00021,TIGR00037,TIGR00042,TIGR00064,TIGR00111,TIGR00134,TIGR00240,TIGR00264,TIGR00270,TIGR00279,TIGR00283,TIGR00291,TIGR00293,TIGR00307,TIGR00308,TIGR00323,TIGR00324,TIGR00335,TIGR00336,TIGR00337,TIGR00373,TIGR00389,TIGR00392,TIGR00398,TIGR00405,TIGR00408,TIGR00422,TIGR00425,TIGR00432,TIGR00442,TIGR00448,TIGR00456,TIGR00458,TIGR00463,TIGR00468,TIGR00471,TIGR00490,TIGR00491,TIGR00501,TIGR00521,TIGR00522,TIGR00549,TIGR00658,TIGR00670,TIGR00729,TIGR00936,TIGR00982,TIGR01008,TIGR01012,TIGR01018,TIGR01020,TIGR01025,TIGR01028,TIGR01038,TIGR01046,TIGR01052,TIGR01060,TIGR01077,TIGR01080,TIGR01309,TIGR01952,TIGR02076,TIGR02153,TIGR02236,TIGR02258,TIGR02338,TIGR02389,TIGR02390,TIGR02651,TIGR03626,TIGR03628,TIGR03629,TIGR03636,TIGR03653,TIGR03665,TIGR03670,TIGR03671,TIGR03672,TIGR03673,TIGR03674,TIGR03677,TIGR03680,TIGR03683,TIGR03684,TIGR03722 PF01496.14,PF13685.1 TIGR01213,TIGR03627
- Step 3: Aligning genomes (align)
The align step will align all identified markers, determine the most likely domain and output a concatenated MSA.
gtdbtk align --identify_dir /tmp/gtdbtk/identify --out_dir /tmp/gtdbtk/align --cpus 2
[2020-08-03 11:04:04] INFO: GTDB-Tk v1.3.0
[2020-08-03 11:04:04] INFO: gtdbtk align --identify_dir /tmp/gtdbtk/identify --out_dir /tmp/gtdbtk/align --cpus 2
[2020-08-03 11:04:04] INFO: Using GTDB-Tk reference data version r95: /srv/db/gtdbtk/official/release95
[2020-08-03 11:04:04] INFO: Aligning markers in 2 genomes with 2 threads.
[2020-08-03 11:04:04] INFO: Processing 2 genomes identified as archaeal.
[2020-08-03 11:04:04] INFO: Read concatenated alignment for 1,672 GTDB genomes.
==> Aligned 2/2 (100%) genomes [ 1.17s/it, ETA 00:00]
[2020-08-03 11:04:07] INFO: Masking columns of archaeal multiple sequence alignment using canonical mask.
==> Masked 1674/1674 (100%) alignments [656.58it/s, ETA 00:00]
[2020-08-03 11:04:09] INFO: Masked archaeal alignment from 32,675 to 5,124 AAs.
[2020-08-03 11:04:09] INFO: 0 archaeal user genomes have amino acids in <10.0% of columns in filtered MSA.
[2020-08-03 11:04:09] INFO: Creating concatenated alignment for 1,674 archaeal GTDB and user genomes.
[2020-08-03 11:04:09] INFO: Creating concatenated alignment for 2 archaeal user genomes.
[2020-08-03 11:04:09] INFO: Done.
- Results
It is important to pay attention to the output, if a genome had a low number of markers identified it will be excluded from the analysis at this step. A warning will appear if that is the case.
Depending on the domain, a prefixed file of either ar122 or bac120 will appear containing the MSA of the user genomes and the GTDB genomes, or just the user genomes (gtdbtk.ar122.msa.fasta and gtdbtk.ar122.user_msa.fasta respectively.)
ls /tmp/gtdbtk/align
align gtdbtk.ar122.user_msa.fasta identify
gtdbtk.ar122.filtered.tsv gtdbtk.log
gtdbtk.ar122.msa.fasta gtdbtk.warnings.log
- Step 4: Classifying genomes (classify)
分类步骤会将基因组放入参考树中,然后确定其最可能的分类。
gtdbtk classify --genome_dir /tmp/gtdbtk/genomes --align_dir /tmp/gtdbtk/align --out_dir /tmp/gtdbtk/classify -x gz --cpus 2
[2020-08-03 11:04:10] INFO: GTDB-Tk v1.3.0
[2020-08-03 11:04:10] INFO: gtdbtk classify --genome_dir /tmp/gtdbtk/genomes --align_dir /tmp/gtdbtk/align --out_dir /tmp/gtdbtk/classify -x gz --cpus 2
[2020-08-03 11:04:10] INFO: Using GTDB-Tk reference data version r95: /srv/db/gtdbtk/official/release95
[2020-08-03 11:04:10] INFO: Placing 2 archaeal genomes into reference tree with pplacer using 2 cpus (be patient).
==> Step 9 of 9: placing genome 2 of 2 (100.00%).
[2020-08-03 11:05:22] INFO: pplacer version: v1.1.alpha19-0-g807f6f3
[2020-08-03 11:05:22] INFO: Calculating RED values based on reference tree.
[2020-08-03 11:05:22] INFO: Calculating average nucleotide identity using FastANI.
[2020-08-03 11:05:22] INFO: fastANI version: 1.31
==> Processed 4/4 (100%) comparisons [ 2.49it/s, ETA 00:00]
[2020-08-03 11:05:24] INFO: 2 genome(s) have been classified using FastANI and pplacer.
[2020-08-03 11:05:24] INFO: Done.
- Results
The two main files output (one again, depending on their domain) are the summary file, and the reference tree containing those genomes (gtdbtk.ar122.summary.tsv, andgtdbtk.ar122.classify.treerespectively). 基因组的分类显示在summary文件中。
ls /tmp/gtdbtk/classify
classify gtdbtk.ar122.summary.tsv gtdbtk.warnings.log
gtdbtk.ar122.classify.tree gtdbtk.log
批量dRep以及GTDB注释
awk 'NR==1{print "method,"$0}$1!~/genome/{split($1,a,"_");print a[2]","$0}' GTDB_for_matbat2_maxbin2/dRep_out/data_tables/genomeInfo.csv GTDB_for_vamb/dRep_out/data_tables/genomeInfo.csv >Stats/vamb_maxbin2_metabat2_all_bins_genomeInfo.csv
awk -F, 'NR==1{print}$3>=50 && $4<=5 || ($3-5*$4)>=50' vamb_maxbin2_metabat2_all_bins_genomeInfo.csv >vamb_maxbin2_metabat2_Hight_quality_bins_genomeInfo.csv
注意
FAQ
- Validating species assignments with average nucleotide identity
GTDB-Tk uses FastANI to estimate the ANI between genomes. We recommend you have FastANI >= 1.32 as this version introduces a fix that makes the results deterministic. A query genome is only classified as belonging to the same species as a reference genome if the ANI between the genomes is within the species ANI circumscription radius (typically, 95%) and the alignment fraction (AF) is >=0.65. In some circumstances, the phylogenetic placement of a query genome may not support the species assignment. GTDB r89 strictly uses ANI to circumscribe species and GTDB-Tk follows this methodology. The species-specific ANI circumscription radii are available from the GTDB website.
报错
由于基因组存在重复id导致Pfam报错
Pfam Search step is stopping with the following error ( Sequence identifiers must be unique. Your fasta file contains two sequences with the same id (Ga0456356_000001_1) ).
Looking at your input file , it seems that every header and sequence is duplicated in the file ( This assembly file has 2 copies of the original assembly).
grep '>' 3300049427_3.fna | sort -n | uniq -c | sort -n
2 >Ga0456356_000001
2 >Ga0456356_000002
2 >Ga0456356_000003
......
解决:
for i in *fa;do printf "${i}\t";grep -F ">" $i|sort |uniq -d ;echo;done |awk 'NF==2' >../all_err_id.stats &
查看已经安装的数据库
(gtdbtk) [yutao@myosin release202]$ gtdbtk check_install # Checking integrity of reference package:即为数据库路径
# 物种名和GCA号对应表
(gtdbtk) [yutao@myosin release202]$ ls $PWD/taxonomy/gtdb_taxonomy.tsv
/share/pasteur/yutao/DATABASE/GTDB_R202/release202/taxonomy/gtdb_taxonomy.tsv
(gtdbtk) [yutao@myosin release202]$ head -n3 $PWD/taxonomy/gtdb_taxonomy.tsv
RS_GCF_000970205.1 d__Archaea;p__Halobacteriota;c__Methanosarcinia;o__Methanosarcinales;f__Methanosarcinaceae;g__Methanosarcina;s__Methanosarcina mazei
RS_GCF_000012285.1 d__Archaea;p__Thermoproteota;c__Thermoproteia;o__Sulfolobales;f__Sulfolobaceae;g__Sulfolobus;s__Sulfolobus acidocaldarius
GB_GCA_003162175.1 d__Archaea;p__Halobacteriota;c__Bog-38;o__Bog-38;f__Bog-38;g__Bog-38;s__Bog-38 sp003162175
# 所有reference的位置
(gtdbtk) [yutao@myosin Two_new_classes]$ find /share/pasteur/yutao/DATABASE/GTDB_R202/release202/fastani/database/ -type f -iname "*.gz" >/share/pasteur/yutao/DATABASE/GTDB_R202/release202/47894_GTDB_GCA_path.tsv
- 例如这里提取所有Eisenbacteria和Krumholzibacteriota门的基因组
(gtdbtk) [yutao@myosin Two_new_classes]$ awk '$2~/p__Eisenbacteria/||$2~/p__Krumholzibacteriota/' /share/pasteur/yutao/DATABASE/GTDB_R202/release202/taxonomy/gtdb_taxonomy.tsv >31gtdb.tsv
# 提取基因组
(reverse-i-search)`fin': find /share/pasteur/yutao/DATABASE/GTDB_R202/release202/fastani/database/ -type f -iname "*.gz" >/share/pasteur/yutao/DATABASE/GTDB_R202/release202/47894_GTDB_GCA_path.tsv
(reverse-i-search)`grep': grep -f 15Krumholzibacteriota_gca.id /share/pasteur/yutao/DATABASE/GTDB_R202/release202/47894_GTDB_GCA_path.tsv|while read i;do ln -s $i Krumholzibacteriota/;done
iTOL可视化问题
- gtdb生成的newick格式的树直接导入iTOL可能丢失叶子节点的问题,是因为叶子的名称包含特殊newick使用的标志字符”;“
gtdb-tk导出的.tree文件即是nwk格式,但是有标注含有“;”,需将其改为其他符号,因为在nwk中,分号“;”认为是进化树结束的标志,即末尾含有一个“;”。并且,把单引号内clade标注前面的数值和冒号删除。之后可以在iTol可视化。
notepad++正则表达式替换如下:
‘[0,1].([0-9]{0,5}):替换为’,删除bootstrap
或者
:[p,k,c,o,f,g,s].*?'和’去除都去除,删除inter node
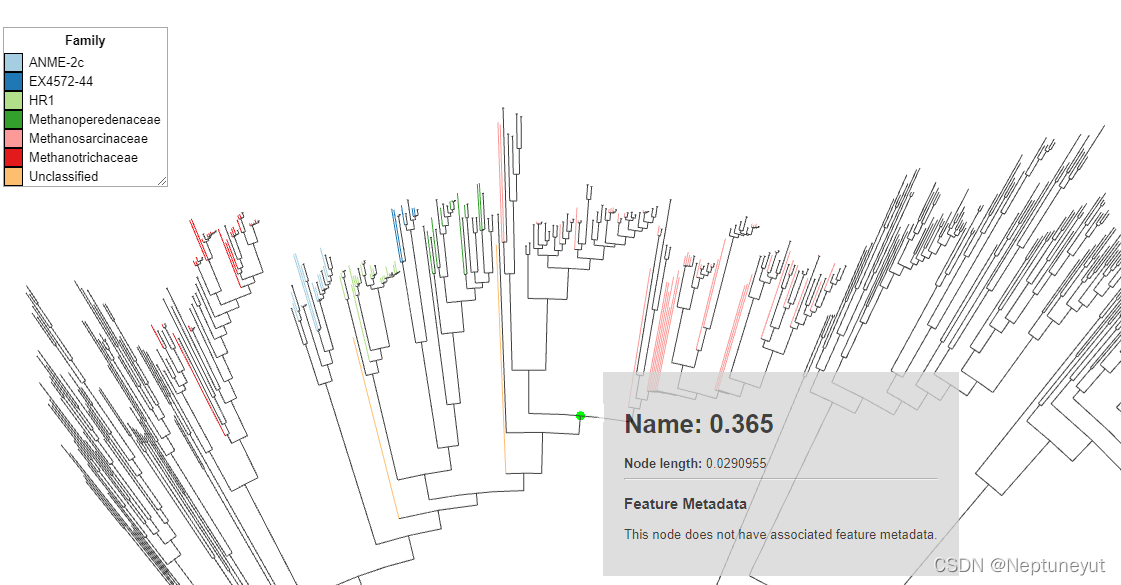
pplacer只是插入树的问题
gtdb的序列,都是在clade的根部。这是典型的pplacer插入的结果。这个结果只能参考看看。所有的树都要从头构建。2000多的序列,也不会花费太长时间。
问题
- 由于基因组存在重复id导致Pfam报错
Pfam Search step is stopping with the following error ( Sequence identifiers must be unique. Your fasta file contains two sequences with the same id (Ga0456356_000001_1) ).
Looking at your input file , it seems that every header and sequence is duplicated in the file ( This assembly file has 2 copies of the original assembly).
grep '>' 3300049427_3.fna | sort -n | uniq -c | sort -n
2 >Ga0456356_000001
2 >Ga0456356_000002
2 >Ga0456356_000003
......
解决:
for i in *fa;do printf "${i}\t";grep -F ">" $i|sort |uniq -d ;echo;done |awk 'NR==2' >../all_err_id.stats &
参考
gtdbtk








![[EFI]戴尔Latitude 5310电脑 Hackintosh 黑苹果efi引导文件](https://img-blog.csdnimg.cn/b50f38253fad460dbf62432e87d7c769.png)


